
Catherine Drennan, a professor of chemistry and biology, likes to wax poetic about the complex chemistry of microbes. “I think they’re elegant and beautiful,” she says. Of course, she also sees their practical applications. “I love the fact that these microbes can be used for bioremediation in hard-to-reach polluted spots,” she says. “Normally, cleaning these delicate environments would upset the balance of living organisms there, but microbes offer a natural way to clean things utilizing chemistry.”
Drennan and Michael Funk, a graduate student in chemistry, are part of a team testing the nuances of how microbes break down hydrocarbons while metabolizing them for growth and survival. Because hydrocarbons are a natural (and major) molecular component of crude oil, microbes could aid cleanup efforts following oil spills and other instances of pollution — as long as the microbes’ hydrocarbon metabolism process proceeds smoothly. The first step in that process calls for the microbe to chemically attack the hydrocarbon to generate a more reactive molecule, but some hydrocarbons are challenging to get a handle on.
To learn more about how to circumvent the inert nature of hydrocarbons, Drennan, Funk, and fellow researchers explored the structural basis of the process by which one microbe — the denitrifying bacterium Thauera aromatica — activated and broke down the hydrocarbon toluene.
The details of their findings, which involve the mapping of previously unknown enzymatic structures, were released in a paper this summer in The Journal of Biological Chemistry.
“When you’re thinking about how an organism breaks a carbon source down and then uses that to make energy for itself,” Drennan says, “you think it’s going to take it and pull it apart, but in this case, it makes a bigger molecule first. That’s because the thing that it wants to add into its metabolism and make energy from is so nonreactive that it has to come up with some way to get a handle on it and make it useful.”
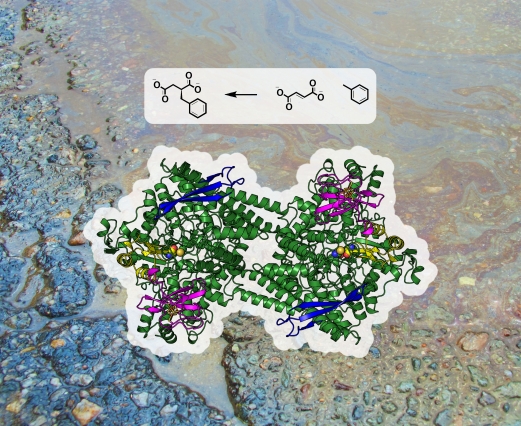
The enzyme benzylsuccinate synthase captures and attaches to the hydrocarbon toluene, a major component in gasoline, to break it down into benzylsuccinate. Shown here are the enzymatic reaction and chemical structures of toluene (right), fumarate (center), and benzylsuccinate (left) against the backdrop of an oil spill. Photo: Michael Funk and Catherine Drennan
Winning combination
The winning combination that enables the breakdown process is the pairing of the enzyme benzylsuccinate synthase (naturally occurring within the microbe) with the substrate molecule fumarate (also present in the microbe) in the presence of toluene. The benzylsuccinate synthase acts as a catalyst, encouraging a reaction between the toluene and the fumarate to generate benzylsuccinate. The conversion of toluene to benzylsuccinate makes it more reactive — as Drennan described above — and the first step in the metabolism process is then possible.

Graduate student Michael Funk working in an anaerobic chamber in the Drennan Lab. Photo: Courtesy of the Drennan Lab
Funk recalls the multi-step process of determining how this enzyme enables the reaction: “A couple of years ago our research revealed the structures of the enzyme benzylsuccinate synthase without anything bound, so that was the first step, to see the overall architecture of the active site. But we still didn’t know how that enzyme would bind to a substrate.”
Funk and Drennan were able to get the substrate fumarate to bind to the enzyme: Fumarate has two carboxyl groups and fits perfectly with the active site. With those two pieces in place, they experimented with different forms of the enzyme until they were able to bind toluene.
Discoveries like this one — which shed light on how hydrocarbons like toluene can be broken down, despite their nonpolar (and therefore usually non-binding) nature — are a critical starting point for more organized efforts in bioremediation (waste management using organisms), Funk says. He is quick to point out, however, that we are still far from any kind of panacea.
“One of the things I didn’t anticipate is how precise the active sites on molecules are,” Funk says. “This enzyme [benzylsuccinate synthase] utilizes toluene, and everything is packed very tightly around that molecule. So it’s not like it can just take anything — it’s very specific for that particular molecule.”
Bioremediation and beyond
This has larger implications for the practice of bioremediation, Funk says: “If you’re going to be doing bioremediation, you’re going to need many different kinds of enzymes that are capable of degrading all different kinds of substrates.”
It’s not just in labs where microbes thrive on hydrocarbons. “Some microbes will grow abundantly near oil refineries, to the point that the oil companies are not happy about it because it’s clogging up all of their pipes,” Drennan says. “This actually gives weight to the idea that microbial bioremediation could work, because it’s evidence that microbes can be very happy and take care of things in such an environment.”
It’s also evidence that the discoveries concerning microbial structure that Drennan and Funk reveal in their paper could have more than one use. “[Pipe erosion] has been an issue because organisms reduce sulfate to hydrogen sulfide,” Funk says, “which is a really nasty way to corrode pipes.”
And pipe erosion in an oil refinery can only mean one thing: heightened probability of a spill.
With this issue in mind, Drennan and Funk acknowledge the irony that their structural discoveries may sooner be used for the inhibition of microbes than for fostering their growth. “I hope that this work doesn’t just lead to an inhibitor that kills the organisms so they don’t corrode the pipes,” Drennan says. “But if it prevents spills, that’s good. You want the right amount of microbes cleaning up the environment, but not so much that they actually pollute it.”
Whatever the immediate uses of their latest discovery, Drennan and Funk’s love affair with microbes is far from over. The DNA sequencing of microbes in particular presents an exciting frontier. “If you have these DNA sequences,” Funk says, “you can turn out a profile of what the microbes are able to degrade in a particular spill situation, as well as what’s been able to slip through because it doesn’t have a degradation pathway. Sequence information can tell you a lot — it can give you a much bigger picture about a whole community of organisms.”